
LIBERTY
Leaf Incorporating Biochemistry Exhibiting Reflectance and Transmittance Yields


|
|
| Introduction
LIBERTY is a general purpose radiative tranfer model for predicting the reflectance and transmittance spectra of a leaf, or stack of leaves in the visible and near infrared wavelengths (400 - 2500 nm). By treating a leaf as an aggregation of cells, with multiple radiation scattering between cells, output spectra is a function of three structural parameters and the combined absorption coefficients of leaf biochemicals. It is simple to use with the user being prompted for input values and the model output is written to an external file for use with graphing and spreadsheet packages as well as for coupling with vegetation canopy or ecosystem models. The code is in C and has been incorporated into the 5-Scale model. |
|
LIBERTY for Windows can use external data files instead of the defaults. This allows the user to easily provide new absorption coefficients: PIGMENT.DAT
ALBINO.DAT
WATER.DAT
LIGCELL.DAT
PROTEIN.DAT
|
|
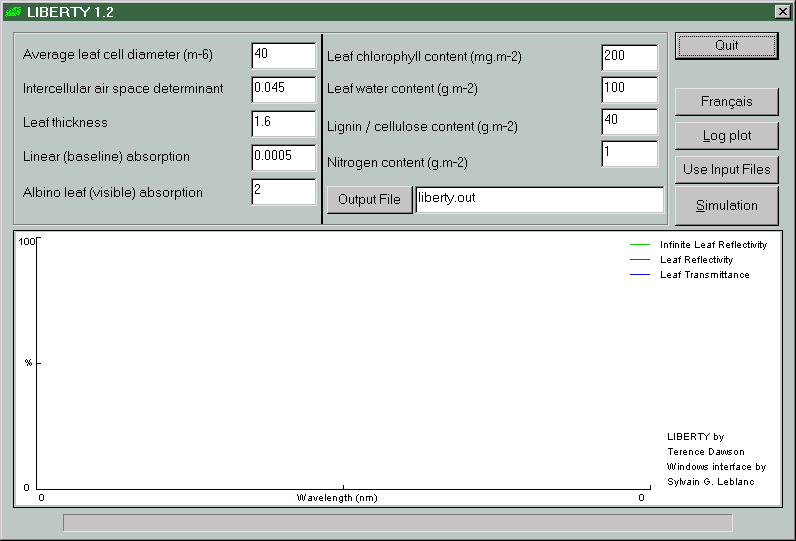
Model inputs |
|
|
|
(and range) |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Dry: 0.0004 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Model Output
|
|
LIBERTY produces a space delimited text file, example as below: |
| W R refl trans
400 0.029629 0.029592 0.035535 405 0.030031 0.029991 0.036120 ... 2500 0.517565 0.414745 0.394991 |
| The first column is the wavelength, second column is the predicted spectra for infinite reflectance as demonstrated by stacked leaves, and the third and fourth column is the predicted single leaf reflectance and transmittance spectra, respectively. |
|
Notes LIBERTY has been developed specifically for conifer needles so you will find the facility to estimate single leaf reflectance and transmittance spectra from stacked needles particularly useful. However, the program is equally valid for any leaf species with a minor caveat; The in-vivo absorption coefficients used were determined from empirical work on various pine species so you may need to re-calibrate variable values for other vegetation types. |